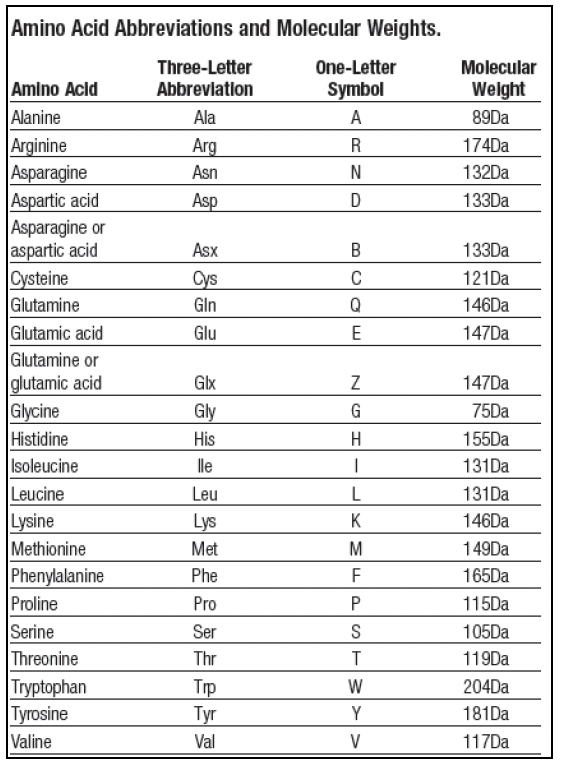
Protein structure, amino acid composition and sequence determine proteome vulnerability to oxidation‐induced damage | The EMBO Journal

Molecules | Free Full-Text | PepFun: Open Source Protocols for Peptide-Related Computational Analysis

Overview of the IPC 2.0 architecture. The input (amino acid sequence in... | Download Scientific Diagram
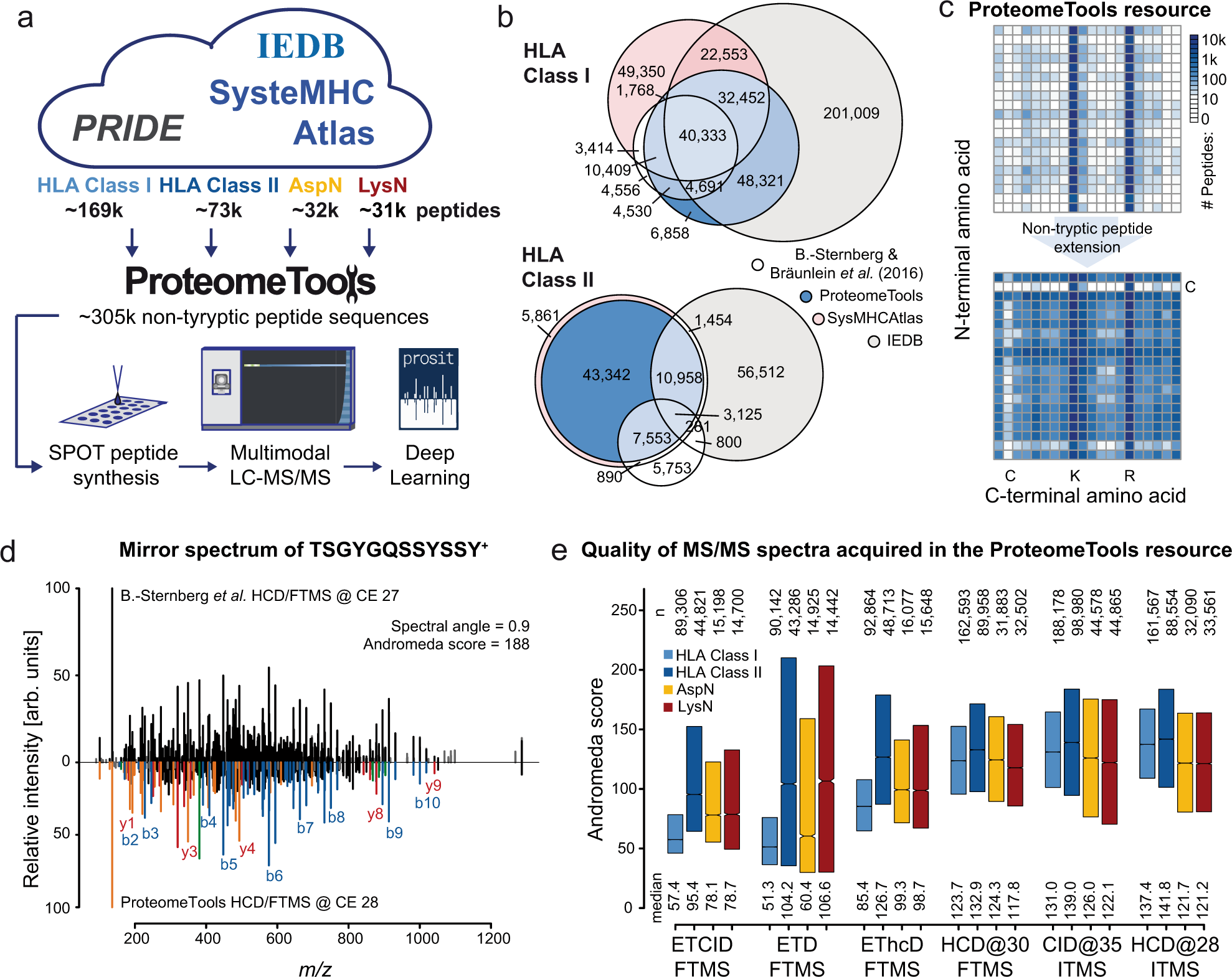
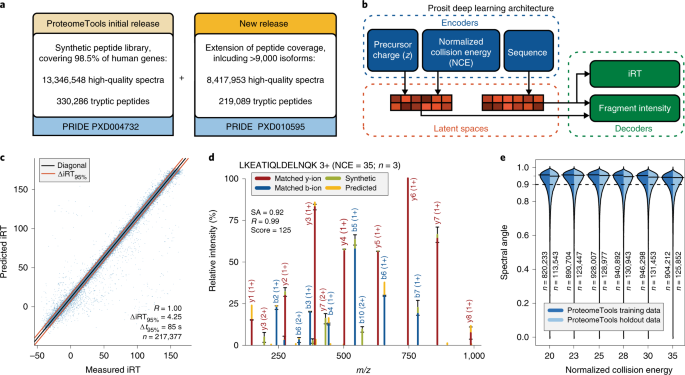
Deep learning boosts sensitivity of mass spectrometry-based immunopeptidomics | Nature Communications
ProteoClade: A taxonomic toolkit for multi-species and metaproteomic analysis | PLOS Computational Biology

pIChemiSt ─ Free Tool for the Calculation of Isoelectric Points of Modified Peptides | Journal of Chemical Information and Modeling
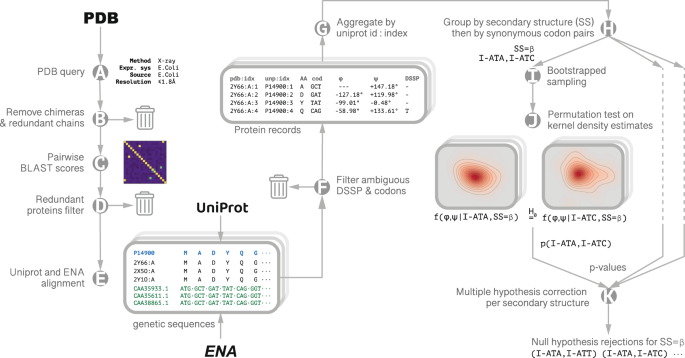
Codon-specific Ramachandran plots show amino acid backbone conformation depends on identity of the translated codon | Nature Communications
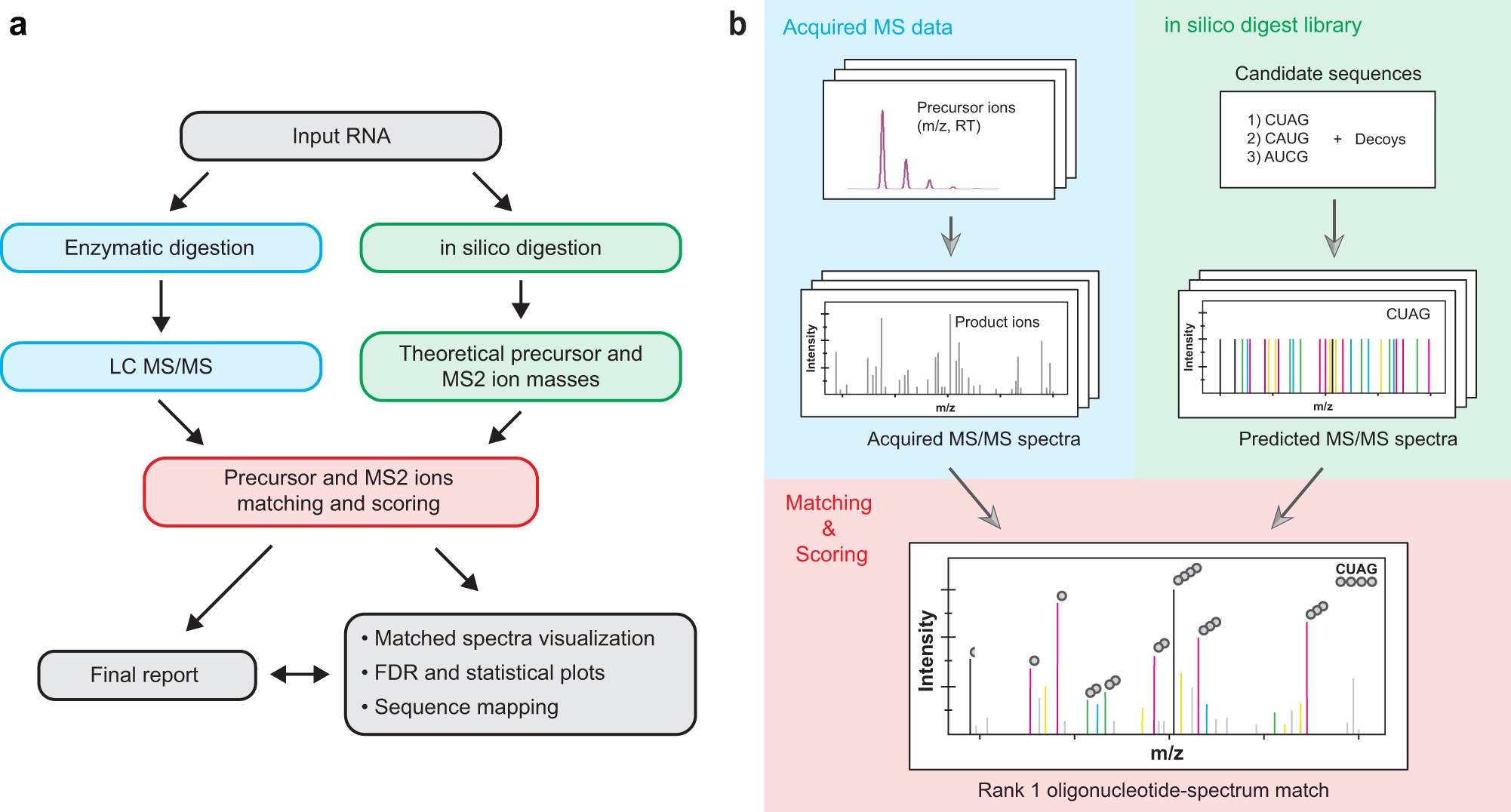
Pytheas: a software package for the automated analysis of RNA sequences and modifications via tandem mass spectrometry | Nature Communications

python - Biopython: How to avoid particular amino acid sequences from a protein so as to plot Ramachandran plot? - Stack Overflow